GROMACS topology for retinal
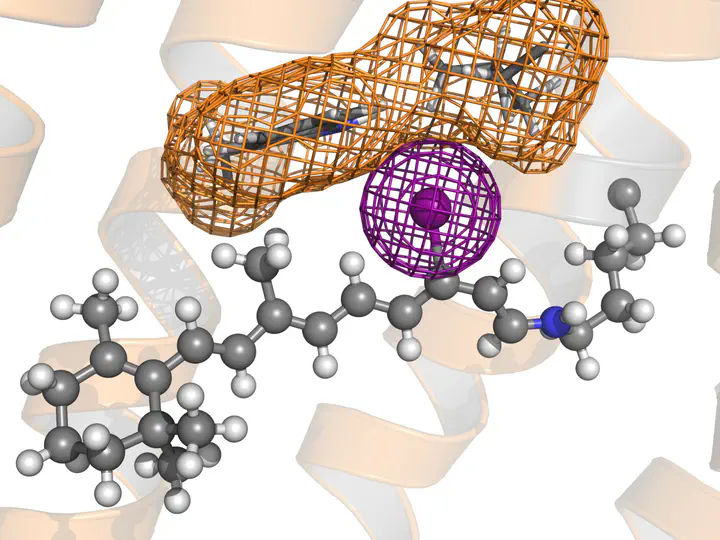
Gromacs topology of a retinal bound to lysine via protonated Schiff base
Here you find the files required to use gmx pdb2gmx to generate a GROMACS topology of retinal bound to lysine via a protonated Schiff base.
These files generate a GROMACS topology of a retinal attached to a lysine, for a simulation using the AMBER99SB*-ILDN force field. Hydrogens may or may not be described as virtual sites. The retinal is in the all-trans state. Note: The image shows an iodo-retinal.
These files come without any warranty of any kind. Check the topology carefully before using it. If you find a bug, please report it to Jochen Hub.
The retinal parameters come originally from Hayashi et al. (see below) and were converted to Gromacs format by Jochen Hub. The AMBER99SB*-ILDN files were take from the Gromacs user contributions in November 2012. If you use these files for publication, please cite (in addition to the AMBER references):