SAXS-Restrained Ensemble Simulations of Intrinsically Disordered Proteins with Commitment to the Principle of Maximum Entropy
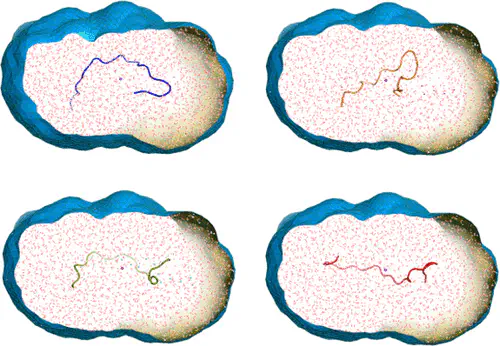
Abstract
Intrinsically disordered proteins (IDPs) play key roles in biology and disease, rationalizing the wide interest in deriving accurate solution ensembles of IDPs. Molecular dynamics (MD) simulations of IDPs often suffer from force-field inaccuracies, suggesting that simulations must be complemented by experimental data to obtain physically correct ensembles. We present a method for integrating small-angle X-ray scattering (SAXS) data on-the-fly into MD simulations of disordered systems, with the aim to bias the conformational sampling toward agreement with ensemble-averaged SAXS data. By coupling a set of parallel replicas to the data and following the principle of maximum entropy, this method applies only a minimal bias. Using the RS peptide as a test case, we analyze the influence of (i) the number of parallel replicas, (ii) the scaling of the force constant for the SAXS-derived biasing energy with the number of parallel replicas, and (iii) the force field. The refined ensembles are cross-validated against experimental 3JHN-Hα couplings and further compared in terms of Cα distance maps and secondary structure content. Remarkably, we find that the applied force field only has a small influence on the SAXS-refined ensemble, suggesting that incorporating SAXS data into MD simulations may greatly reduce the force-field bias.