Computational Biophysics Group
Welcome to the Computational Biophysics Group at Saarland University.
We develop methods related to molecular dynamics simulations, with the aim to understand the relationship between structure, dynamics, and function of biological macromolecules.
Temporary contact: Aouregan Lemée
We have several interesting Bachelor and Master projects available. Find out more.
We do not currently have any open positions. However, we are always interested in hearing from ambitious PhD candidates or postdocs who are willing to apply for their own funding. Find out more.
Research Topics
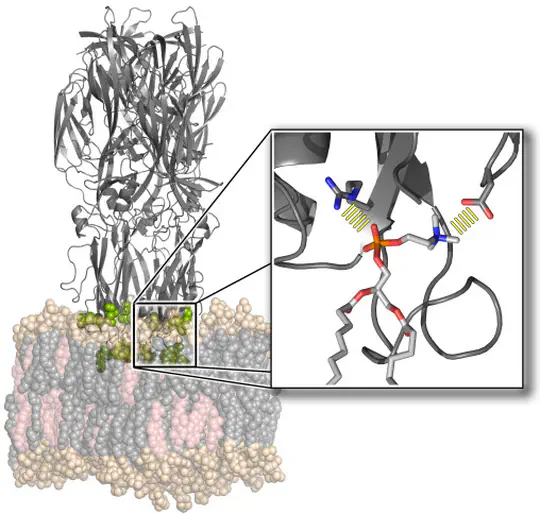
The function of biological membranes goes far beyond the formation of a mere barrier. Membranes are subject to ongoing structural remodeling, which is controlled by interactions with proteins and by the lipid composition. We develop free energy calculation techniques to understand how membrane composition and interactions with proteins (such as viral fusion proteins) enable functionally important events at membranes including membrane fusion, pore formation, or drug permeation.
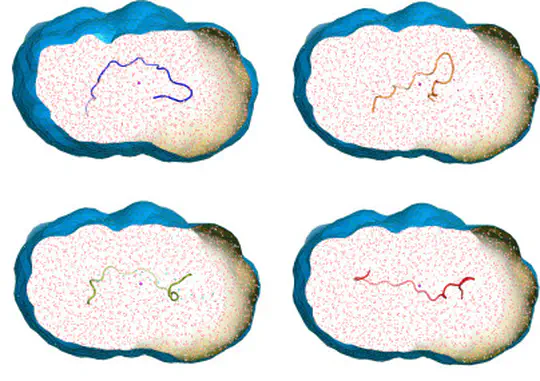
Collecting experimental data is often difficult – but the interpretation of the data may be even more challenging, for instance because the information content of the experimental signals is low. We develop methods for combining MD simulations with experimental data to get the best of two worlds, with some focus on small-angle X-ray and neutron scattering data (SAXS/SANS). Our developments involve accurate SAXS/SANS predictions, protein structure and ensemble refinement, studies on the protein hydration shell, and modeling of experiments at X-ray free electron lasers. We share our methods via the web server WAXSiS and GROMACS-SWAXS.
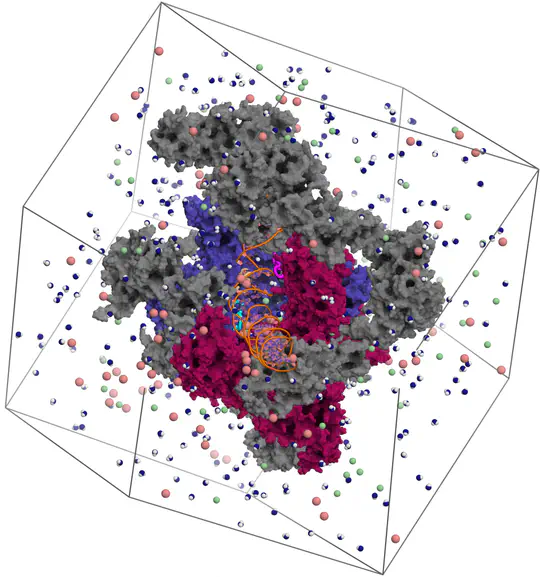
Proteins are not static building blocks but instead carry out their function –and malfunction– by structural transitions (Structure-function-dynamics relationship). We combine MD simulations with experiential data and enhanced-sampling techniques, to observe proteins while they function in atomic detail. Our portfolio comprises studies of molecular motors, protein-RNA/DNA complexes, membrane channels, and enzymes related to cancer progression.
Latest Publications
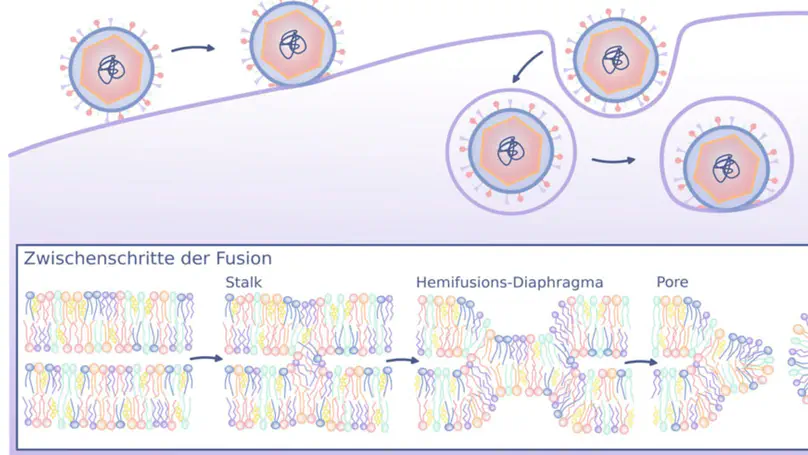
Biological membranes display remarkable compositional diversity with functional consequences. In viral infections, lipids play active roles beyond forming a structural barrier: they modulate fusion protein binding, and tune membrane mechanics to prime them for efficient fusion, while viruses hijack host lipid metabolism to support replication. This highlights how membrane remodeling is exploited throughout the viral life cycle and points to potential avenues for antiviral intervention.
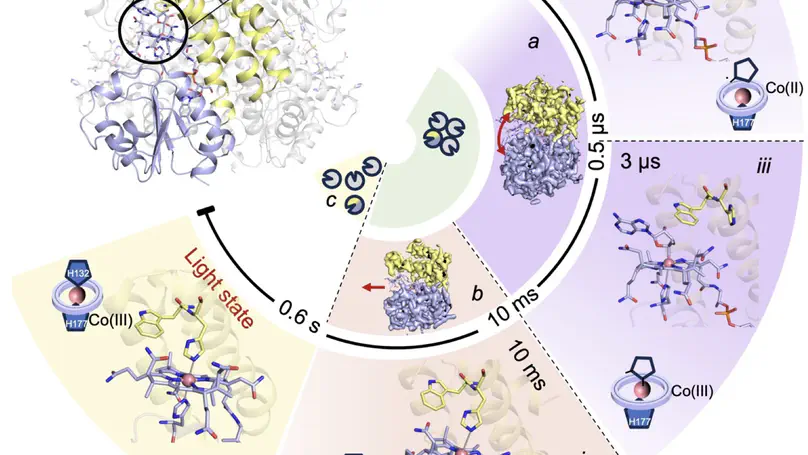
Photoreceptor proteins initiate, regulate and control fundamental biological processes such as vision, photosynthesis and circadian rhythms. A large photoreceptor subfamily uses vitamin B12 derivatives for light sensing, contrasting with the well-established mode of action of these organometallic derivatives in thermally activated enzymatic reactions. The molecular mechanism of B12 photoreception and how this differs to the thermal pathways remain unknown. Here we provide a detailed spatio-temporal description of photoactivation in the prototypical tetrameric B12 photoreceptor CarH from nanoseconds to seconds by using an integrative approach, combining time- and temperature-resolved structural and spectroscopic methods with quantum chemical calculations. High resolution structural snapshots of key intermediates illustrate how photocleavage of a Co–C bond triggers a pathway of structural changes that propagate throughout CarH from the B12 chromophore, via a previously unknown adduct, to finally cause tetramer dissociation. These unique intermediates, which differentiate CarH from thermally-activated B12 enzymes, steer the photoactivation pathway and act as the molecular bridge between photochemical and photobiological timescales. Our results offer a spatio-temporal understanding of CarH photoactivation and pave the way for designing and optimising B12-dependent photoreceptors for future optogenetic applications.
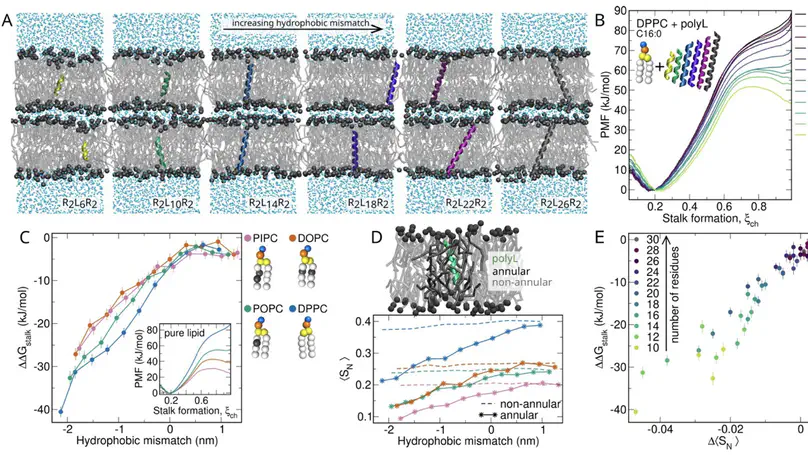
Membrane fusion is a fundamental process involved in exocytosis, fertilization, or cell entry by enveloped viruses. Membrane fusion is facilitated by fusion proteins, which are anchored in membranes by helical transmembrane domains (TMDs). Previous studies showed that TMD variations may alter the fusion efficiency, suggesting that TMDs are not merely passive anchors, however the mechanism by which TMDs drive fusion is not well understood. We used high-throughput coarse-grained molecular dynamics simulations and free energy calculations to quantify effects of TMDs on the formation of the first fusion intermediate, that is, of a fusion stalk. We analyzed five physiologically relevant TMDs derived from viral fusion proteins and the SNARE complex embedded in various lipid environments. We find that the addition of TMDs favors stalk formation by typically 10 to 30 kJ/mol in a concentration-dependent manner. Using helices with sequences R2LnR2 (n=6,…, 26), we find that negative hydrophobic mismatch between the TMD and the membrane core strongly promotes fusion. Analysis of the lipid tail order parameters of annular lipids revealed a strong correlation between stalk stabilization and induced lipid disorder. Together, our findings suggest that TMDs actively contribute to membrane fusogenicity by locally perturbing the membrane order.
Funding
Present and former